|
|
“Experimental reconstructions of 3D atomic structures from electron microscopy images using a Bayesian genetic algorithm”. De Backer A, Van Aert S, Faes C, Arslan Irmak E, Nellist PD, Jones L, N P J Computational Materials 8, 216 (2022). http://doi.org/10.1038/s41524-022-00900-w
Abstract: We introduce a Bayesian genetic algorithm for reconstructing atomic models of monotype crystalline nanoparticles from a single projection using Z-contrast imaging. The number of atoms in a projected atomic column obtained from annular dark field scanning transmission electron microscopy images serves as an input for the initial three-dimensional model. The algorithm minimizes the energy of the structure while utilizing a priori information about the finite precision of the atom-counting results and neighbor-mass relations. The results show promising prospects for obtaining reliable reconstructions of beam-sensitive nanoparticles during dynamical processes from images acquired with sufficiently low incident electron doses.
Keywords: A1 Journal article; Electron microscopy for materials research (EMAT)
DOI: 10.1038/s41524-022-00900-w
|


|
|
|
“Deep convolutional neural networks to restore single-shot electron microscopy images”. Lobato I, Friedrich T, Van Aert S, N P J Computational Materials 10, 10 (2024). http://doi.org/10.1038/s41524-023-01188-0

Abstract: Advanced electron microscopy techniques, including scanning electron microscopes (SEM), scanning transmission electron microscopes (STEM), and transmission electron microscopes (TEM), have revolutionized imaging capabilities. However, achieving high-quality experimental images remains a challenge due to various distortions stemming from the instrumentation and external factors. These distortions, introduced at different stages of imaging, hinder the extraction of reliable quantitative insights. In this paper, we will discuss the main sources of distortion in TEM and S(T)EM images, develop models to describe them, and propose a method to correct these distortions using a convolutional neural network. We validate the effectiveness of our method on a range of simulated and experimental images, demonstrating its ability to significantly enhance the signal-to-noise ratio. This improvement leads to a more reliable extraction of quantitative structural information from the images. In summary, our findings offer a robust framework to enhance the quality of electron microscopy images, which in turn supports progress in structural analysis and quantification in materials science and biology.
Keywords: A1 Journal article; Electron microscopy for materials research (EMAT)
DOI: 10.1038/s41524-023-01188-0
|
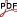


|
|
|
“Investigating lattice strain in Au nanodecahedrons”. Goris B, De Beenhouwer J, de Backer A, Zanaga D, Batenburg J, Sanchez-Iglesias A, Liz-Marzan L, Van Aert S, Sijbers J, Van Tendeloo G, Bals S, , 11 (2016). http://doi.org/10.1002/9783527808465.EMC2016.5519
Keywords: P1 Proceeding; Electron microscopy for materials research (EMAT); Vision lab
DOI: 10.1002/9783527808465.EMC2016.5519
|

|
|
|
“Phase retrieval from 4-dimensional electron diffraction datasets”. Friedrich T, Yu C-P, Verbeek J, Pennycook T, Van Aert S, Proceedings
T2 –, IEEE International Conference on Image Processing (ICIP), SEP 19-22, 2021, Electr. network , 3453 (2021). http://doi.org/10.1109/ICIP42928.2021.9506709

Abstract: We present a computational imaging mode for large scale electron microscopy data, which retrieves a complex wave from noisy/sparse intensity recordings using a deep learning approach and subsequently reconstructs an image of the specimen from the Convolutional Neural Network (CNN) predicted exit waves. We demonstrate that an appropriate forward model in combination with open data frameworks can be used to generate large synthetic datasets for training. In combination with augmenting the data with Poisson noise corresponding to varying dose-values, we effectively eliminate overfitting issues. The U-NET[1] based architecture of the CNN is adapted to the task at hand and performs well while maintaining a relatively small size and fast performance. The validity of the approach is confirmed by comparing the reconstruction to well-established methods using simulated, as well as real electron microscopy data. The proposed method is shown to be effective particularly in the low dose range, evident by strong suppression of noise, good spatial resolution, and sensitivity to different atom types, enabling the simultaneous visualisation of light and heavy elements and making different atomic species distinguishable. Since the method acts on a very local scale and is comparatively fast it bears the potential to be used for near-real-time reconstruction during data acquisition.
Keywords: P1 Proceeding; Engineering sciences. Technology; Electron microscopy for materials research (EMAT)
DOI: 10.1109/ICIP42928.2021.9506709
|
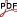


|
|
|
Zhang Z, Lobato I, Brown H, Jannis D, Verbeeck J, Van Aert S, Nellist P (2023) Generalised oscillator strength for core-shell electron excitation by fast electrons based on Dirac solutions

Abstract: Inelastic excitation as exploited in Electron Energy Loss Spectroscopy (EELS) contains a rich source of information that is revealed in the scattering process. To accurately quantify core-loss EELS, it is common practice to fit the observed spectrum with scattering cross-sections calculated using experimental parameters and a Generalized Oscillator Strength (GOS) database [1]. The GOS is computed using Fermi’s Golden Rule and orbitals of bound and excited states. Previously, the GOS was based on Hartree-Fock solutions [2], but more recently Density Functional Theory (DFT) has been used [3]. In this work, we have chosen to use the Dirac equation to incorporate relativistic effects and have performed calculations using Flexible Atomic Code (FAC) [4]. This repository contains a tabulated GOS database based on Dirac solutions for computing double differential cross-sections under experimental conditions. We hope the Dirac-based GOS database can benefit the EELS community for both academic use and industry integration. Database Details: – Covers all elements (Z: 1-108) and all edges – Large energy range: 0.01 – 4000 eV – Large momentum range: 0.05 -50 Å-1 – Fine log sampling: 128 points for energy and 256 points for momentum – Data format: GOSH [3] Calculation Details: – Single atoms only; solid-state effects are not considered – Unoccupied states before continuum states of ionization are not considered; no fine structure – Plane Wave Born Approximation – Frozen Core Approximation is employed; electrostatic potential remains unchanged for orthogonal states when – core-shell electron is excited – Self-consistent Dirac–Fock–Slater iteration is used for Dirac calculations; Local Density Approximation is assumed for electron exchange interactions; continuum states are normalized against asymptotic form at large distances – Both large and small component contributions of Dirac solutions are included in GOS – Final state contributions are included until the contribution of the previous three states falls below 0.1%. A convergence log is provided for reference. Version 1.1 release note: – Update to be consistent with GOSH data format [3], all the edges are now within a single hdf5 file. A notable change in particular, the sampling in momentum is in 1/m, instead of previously in 1/Å. Great thanks to Gulio Guzzinati for his suggestions and sending conversion script. Version 1.2 release note: – Add “File Type / File version” information [1] Verbeeck, J., and S. Van Aert. Ultramicroscopy 101.2-4 (2004): 207-224. [2] Leapman, R. D., P. Rez, and D. F. Mayers. The Journal of Chemical Physics 72.2 (1980): 1232-1243. [3] Segger, L, Guzzinati, G, & Kohl, H. Zenodo (2023). doi:10.5281/zenodo.7645765 [4] Gu, M. F. Canadian Journal of Physics 86(5) (2008): 675-689.
Keywords: Dataset; Electron microscopy for materials research (EMAT)
DOI: 10.5281/ZENODO.8360240
|

|
|
|
“Low-dose 4D-STEM tomography for beam-sensitive nanocomposites”. Hugenschmidt M, Jannis D, Kadu AA, Grünewald L, De Marchi S, Perez-Juste J, Verbeeck J, Van Aert S, Bals S, ACS materials letters 6, 165 (2023). http://doi.org/10.1021/ACSMATERIALSLETT.3C01042

Abstract: Electron tomography is essential for investigating the three-dimensional (3D) structure of nanomaterials. However, many of these materials, such as metal-organic frameworks (MOFs), are extremely sensitive to electron radiation, making it difficult to acquire a series of projection images for electron tomography without inducing electron-beam damage. Another significant challenge is the high contrast in high-angle annular dark field scanning transmission electron microscopy that can be expected for nanocomposites composed of a metal nanoparticle and an MOF. This strong contrast leads to so-called metal artifacts in the 3D reconstruction. To overcome these limitations, we here present low-dose electron tomography based on four-dimensional scanning transmission electron microscopy (4D-STEM) data sets, collected using an ultrafast and highly sensitive direct electron detector. As a proof of concept, we demonstrate the applicability of the method for an Au nanostar embedded in a ZIF-8 MOF, which is of great interest for applications in various fields, including drug delivery.
Keywords: A1 Journal article; Electron microscopy for materials research (EMAT)
DOI: 10.1021/ACSMATERIALSLETT.3C01042
|
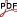

|
|
|
Grü,newald L, Chezganov D, De Meyer R, Orekhov A, Van Aert S, Bogaerts A, Bals S, Verbeeck J (2023) Supplementary Information for “In-situ Plasma Studies using a Direct Current Microplasma in a Scanning Electron Microscope”
Abstract: Supplementary information for the article “In-situ Plasma Studies using a Direct Current Microplasma in a Scanning Electron Microscope” containing the videos of in-situ SEM imaging (mp4 files), raw data/images, and Jupyter notebooks (ipynb files) for data treatment and plots. Link to the preprint: https://doi.org/10.48550/arXiv.2308.15123 Explanation of the data files can be found in the Information.pdf file. The Videos folder contains the in-situ SEM image series mentioned in the paper. If there are any questions/bugs, feel free to contact me at lukas.grunewaldatuantwerpen.be
Keywords: Dataset; Engineering sciences. Technology; Electron microscopy for materials research (EMAT); Plasma Lab for Applications in Sustainability and Medicine – Antwerp (PLASMANT)
DOI: 10.5281/ZENODO.8042030
|

|
|
|
Cioni M, Delle Piane M, Polino D, Rapetti D, Crippa M, Arslan Irmak E, Pavan GM, Van Aert S, Bals S (2024) Data for Sampling Real‐Time Atomic Dynamics in Metal Nanoparticles by Combining Experiments, Simulations, and Machine Learning

Abstract: Even at low temperatures, metal nanoparticles (NPs) possess atomic dynamics that are key for their properties but challenging to elucidate. Recent experimental advances allow obtaining atomic‐resolution snapshots of the NPs in realistic regimes, but data acquisition limitations hinder the experimental reconstruction of the atomic dynamics present within them. Molecular simulations have the advantage that these allow directly tracking the motion of atoms over time. However, these typically start from ideal/perfect NP structures and, suffering from sampling limits, provide results that are often dependent on the initial/putative structure and remain purely indicative. Here, by combining state‐of‐the‐art experimental and computational approaches, how it is possible to tackle the limitations of both approaches and resolve the atomistic dynamics present in metal NPs in realistic conditions is demonstrated. Annular dark‐field scanning transmission electron microscopy enables the acquisition of ten high‐resolution images of an Au NP at intervals of 0.6 s. These are used to reconstruct atomistic 3D models of the real NP used to run ten independent molecular dynamics simulations. Machine learning analyses of the simulation trajectories allows resolving the real‐time atomic dynamics present within the NP. This provides a robust combined experimental/computational approach to characterize the structural dynamics of metal NPs in realistic conditions.
Keywords: Dataset; Engineering sciences. Technology; Electron microscopy for materials research (EMAT)
DOI: 10.5281/ZENODO.10997963
|

|

